Updated: 16 Mar 2026
This notebook takes you through a simple CBED simulation for the same crystal used in the thesis and creates a RGB overlay to compare the differences. Feel free to play around with the simulation parameters like the electron energy. This notebook also does a PACBED scan and compares the different models for that as well.
import abtem
import ase
import matplotlib.pyplot as plt
import numpy as npSTO_crystal = ase.Atoms(
"SrTiO3",
scaled_positions=[
(0.0, 0.0, 0.0),
(0.5, 0.5, 0.5),
(0.5, 0.0, 0.5),
(0.5, 0.5, 0.0),
(0.0, 0.5, 0.5),
],
cell=[3.91270131, 3.91270131, 3.91270131, 90, 90, 90],
pbc=True
)
STO_orthorhombic = abtem.orthogonalize_cell(STO_crystal)abtem.show_atoms(STO_orthorhombic,plane='yz',scale=0.5,tight_limits=True,show_periodic=True)(<Figure size 640x480 with 1 Axes>, <Axes: xlabel='y [Å]', ylabel='z [Å]'>)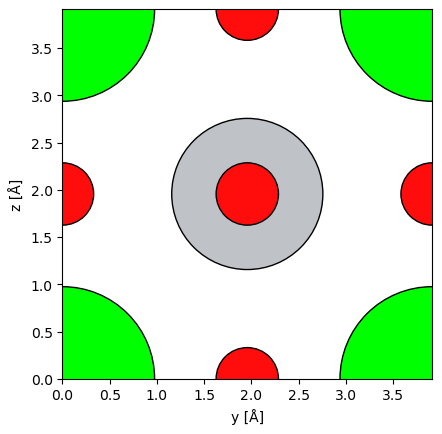
sampling = 0.1 #Angstrom
slice_thickness = 0.5 #Angstrom
thickness = 48 #unit cells
sample_size = (12,12,thickness)
energy = 20e3 #eV
semiangle=20unit_cell_potential = abtem.Potential(
STO_orthorhombic,
sampling=sampling,
parametrization="lobato",
slice_thickness=slice_thickness,
projection="finite",
# device='gpu'
)
potential = abtem.CrystalPotential(
unit_cell_potential,
repetitions=sample_size,
)probe = abtem.Probe(
energy=energy,
semiangle_cutoff=semiangle,
# device='gpu'
).match_grid(potential)
probe.show()Loading...
<abtem.visualize.visualizations.Visualization at 0x7614f8340a10>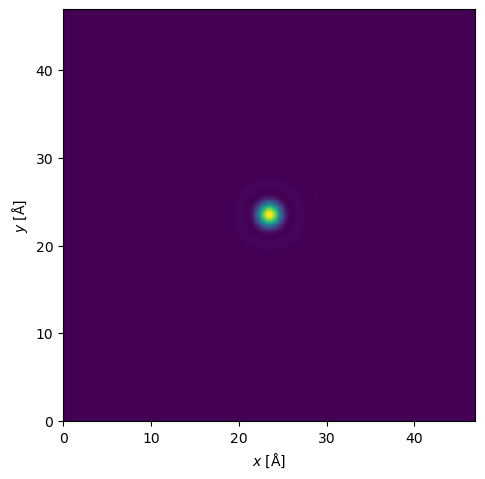
from abtem.multislice import RealSpaceMultislice
detector = abtem.PixelatedDetector(max_angle=None)
CMS = RealSpaceMultislice()
PCMS = RealSpaceMultislice(
order = 3,
)
FCMS = RealSpaceMultislice(
order = 3,
expansion_scope='full'
)exitwave_cms = probe.multislice(potential=potential,detectors=detector, algorithm = CMS,pbar=True,lazy=False)Loading...
exitwave_pcms = probe.multislice(potential=potential,detectors=detector, algorithm = PCMS,pbar=True,lazy=False)Loading...
exitwave_fcms = probe.multislice(potential=potential,detectors=detector, algorithm = FCMS,pbar=True,lazy=False)Loading...
all_exitwaves = abtem.stack(
(
exitwave_cms,
exitwave_pcms,
exitwave_fcms
),
(
'CMS',
'PCMS',
'FCMS'
)
)
all_exitwaves.show(
explode = True,
cbar=True,
cmap='magma',
figsize=(16, 9),
vmin=0,
common_color_scale=True,
power=0.5
)<abtem.visualize.visualizations.Visualization at 0x7614b8cffb90>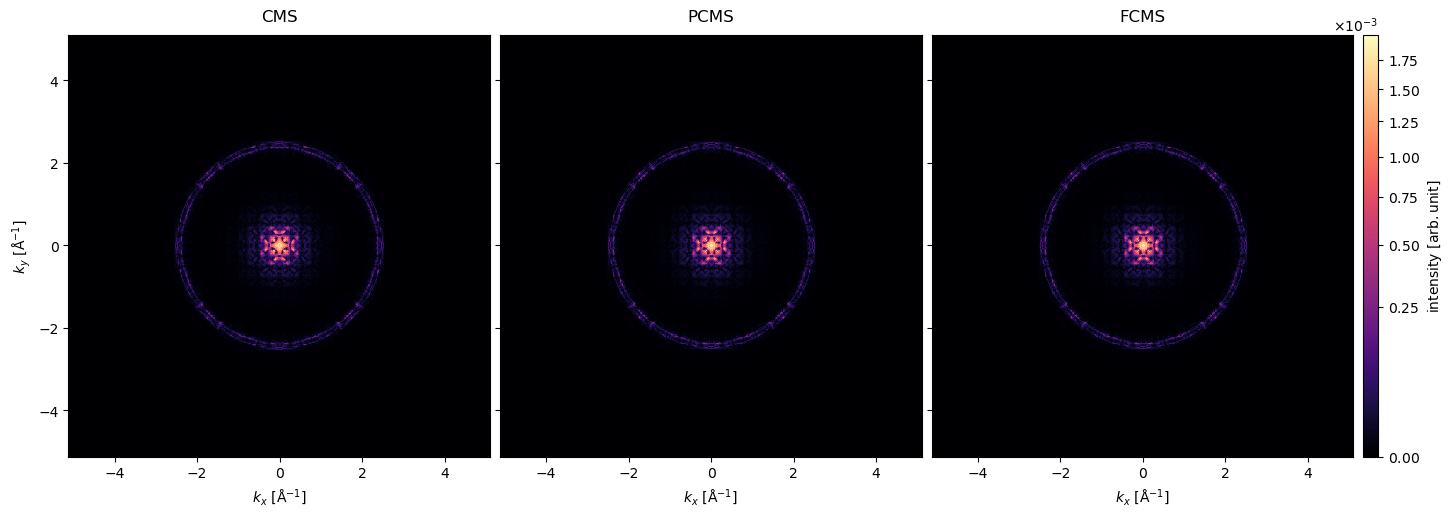
threshold = 0.98
cms_array = exitwave_cms.array
R_quantile = np.quantile(cms_array, threshold)
cms_array = np.clip(cms_array, 0, R_quantile)
R = cms_array / cms_array.max()
pcms_array = exitwave_pcms.array
G_quantile = np.quantile(pcms_array, threshold)
pcms_array = np.clip(pcms_array, 0, G_quantile)
G = pcms_array / pcms_array.max()
fcms_array = exitwave_fcms.array
B_quantile = np.quantile(fcms_array, threshold)
fcms_array = np.clip(fcms_array, 0, B_quantile)
B = fcms_array / fcms_array.max()
rgb_image = np.stack([R, G, B], axis=-1)
plt.imshow(rgb_image, vmin=0)
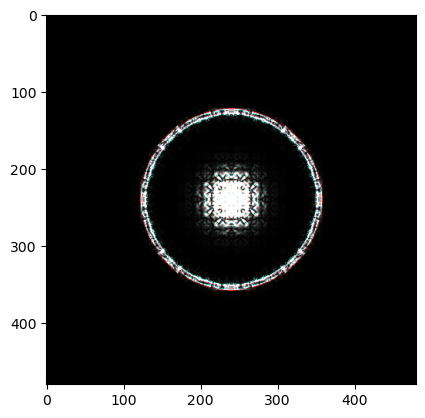
PACBED¶
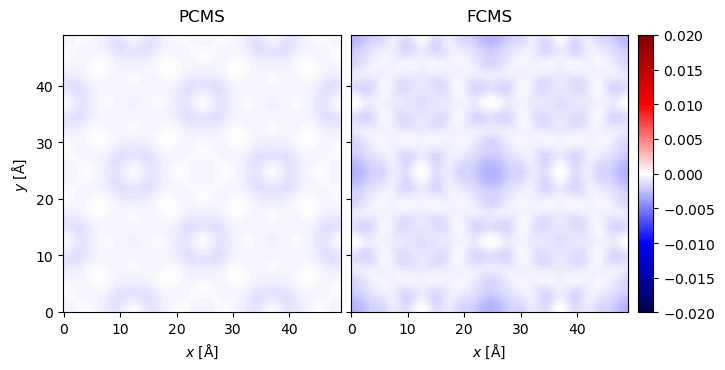
Here we perform a PACBED scan for a higher electron energy and plot the relative difference between the PCMS/FCMS and the CMS as explained in Section. STEM.
haadf = abtem.AnnularDetector(inner=25, outer=None)
probe = abtem.Probe(energy=200e3, semiangle_cutoff=semiangle, device='gpu').match_grid(potential)
grid_scan = abtem.GridScan(
start=[0, 0],
end=(STO_orthorhombic.cell[0,0]*1, STO_orthorhombic.cell[1,1]*1),
sampling=probe.aperture.nyquist_sampling,
potential=potential,
)
fig, ax = abtem.show_atoms(STO_orthorhombic*(12,12,1))
grid_scan.add_to_plot(ax)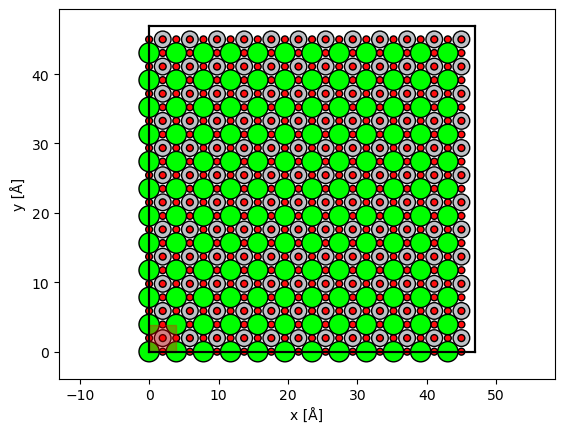
measurements_cms = probe.scan(potential, scan=grid_scan, detectors=haadf, lazy=False, algorithm=CMS, pbar=True)Loading...
measurements_pcms = probe.scan(potential, scan=grid_scan, detectors=haadf, lazy=False, algorithm=PCMS, pbar=True)Loading...
measurements_fcms = probe.scan(potential, scan=grid_scan, detectors=haadf, lazy=False, algorithm=FCMS, pbar=True)Loading...
interpolation_value = 0.05
all_exitwaves = abtem.stack(
(
measurements_cms.tile((2,2)).interpolate(sampling=interpolation_value),
measurements_pcms.tile((2,2)).interpolate(sampling=interpolation_value),
measurements_fcms.tile((2,2)).interpolate(sampling=interpolation_value)
),
(
'CMS',
'PCMS',
'FCMS'
)
)
all_exitwaves.show(
explode = True,
cbar=True,
cmap='inferno',
figsize=(14, 8),
common_color_scale=True,
)<abtem.visualize.visualizations.Visualization at 0x7614b033f4d0>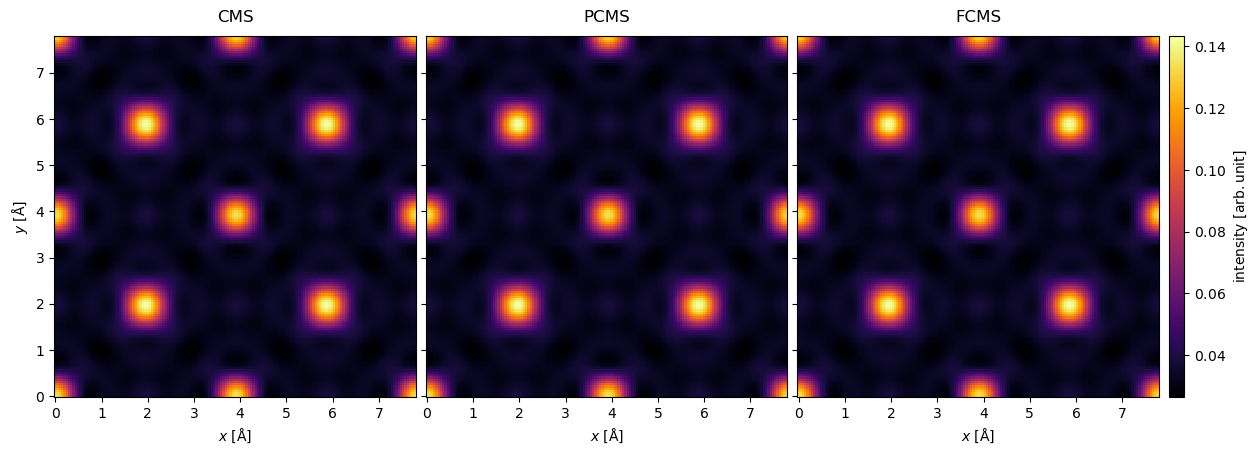
from abtem.measurements import Images
diff_rel_pcms = (measurements_pcms.tile((2,2)).interpolate(sampling=interpolation_value).array - measurements_cms.tile((2,2)).interpolate(sampling=interpolation_value).array) / (1e-6 + measurements_cms.tile((2,2)).interpolate(sampling=interpolation_value).array)
diff_rel_fcms = (measurements_fcms.tile((2,2)).interpolate(sampling=interpolation_value).array - measurements_cms.tile((2,2)).interpolate(sampling=interpolation_value).array) / (1e-6 + measurements_cms.tile((2,2)).interpolate(sampling=interpolation_value).array)
diff_rel_pcms_img = Images(array = diff_rel_pcms, sampling=probe.aperture.nyquist_sampling)
diff_rel_fcms_img = Images(array = diff_rel_fcms, sampling=probe.aperture.nyquist_sampling)
all_measurements = abtem.stack(
(
diff_rel_pcms_img,
diff_rel_fcms_img
),
(
'PCMS',
'FCMS'
)
)
all_measurements.show(
cmap='seismic',
vmin=-0.02,
vmax=0.02,
explode=True,
figsize=(8, 5),
common_color_scale=True,
cbar=True
)<abtem.visualize.visualizations.Visualization at 0x7614991ee250>